The genetic one, that is…
One of the very best things about genealogy is having a genealogy buddy: someone who shares the passion and — if you’re very very lucky — an awful lot of your genes.
The Legal Genealogist is very very lucky. I have a first cousin, Paula — youngest daughter of my mother’s youngest sister — who is my genealogy buddy.
We often go to conferences together, we’ve done road trips together, we share notes on our latest finds.
And, just recently, we had a chance to match up the one type of DNA where she and I have to be the most alike: our mitochondrial DNA (mtDNA).
This is the type of DNA everyone has — so both men and women can be tested. But we all inherit it only from our mothers — so it gives us a look at our mother’s mother’s mother’s line all the way back into distant time.1
Now we are both daughters of daughters of a single woman: our grandmother was Opal (Robertson) Cottrell, born in Texas in 1898 and died in Virginia in 1995.2 Our mtDNA came to us in a direct unbroken line:
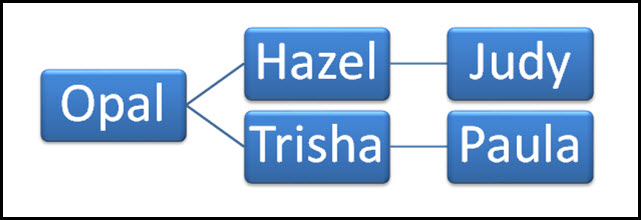
And we both tested all the way out — what’s called the Full Mitochondrial Sequence at Family Tree DNA.
Now, the expectation would be that my mtDNA and Paula’s mtDNA would be identical. We tend to think of genetic distance as the size of the genetic drift that occurs over time — and the genetic distance between maternal first cousins when it comes to mtDNA would have to be essentially zero, now, wouldn’t it?
Not surprisingly, we both share the same mitochondrial haplogroup: we are H3g, described as “a human mitochondrial DNA (mtDNA) haplogroup that likely originated in Southwest Asia” and has a generally Western European distribution with “peak among Basques (13.9%), Galicians (8.3%) and Sardinians (8.5%).”3
And, not surprisingly, at that highest level of testing, looking at the entirety of our mtDNA, we share the same number of matches in the FTDNA database: we both have 15 of these top-level matches.
And, not surprisingly, each of us is on the other’s match list.
And that’s when we do get the surprise: neither of us is the other’s closest match. Each of us has five people — the same five, as a matter of fact — who are closer matches. Five people who are utter total strangers to us, whose mtDNA is closer to our own than our own first cousin.
 That’s because of something called a heteroplasmy. That, by definition, is “the presence of a mixture of more than one type of … mitochondrial DNA … within a cell or individual.”4
That’s because of something called a heteroplasmy. That, by definition, is “the presence of a mixture of more than one type of … mitochondrial DNA … within a cell or individual.”4
I have one in the initial hypervariable region (HVR1), and Paula has one in the coding region. In both cases, the testing system compares our results to that of what’s called a reference sequence. Somewhere back in time, somebody’s entire mtDNA was figured out (sequenced) and everyone else’s now compared to that reference sample.5
In my case, the reference sample shows a value of T in a particular location, but my mtDNA has both T and C. In Paula’s case, the reference sample shows as value of C in a particular location, and she has both T and C.
When we’re compared to those five people, since none of them has the heteroplasmy that I have, each of them is a genetic distance of 1 from me. And none of them has the heteroplasmy that Paula has, so each of them is a genetic distance of 1 from her.
But when Paula and I are compared to each other, the one difference I have that she doesn’t and the one difference she has that I don’t are added together and we’re reported as a genetic distance of 2 from each other.
Get the drift here?
Here’s a case where genetic distance really isn’t an indicator of how closely related two people are. It really is just what it’s defined to be: “the number of differences or mutations between two sets of … mitochondrial DNA test results.”6
Sigh… nobody ever said this would be easy.
SOURCES
Chart: “How do I know if I have an mtDNA (mitochondrial DNA) heteroplasmy? What is the nomenclature?,” Family Tree DNA Learning Center (https://www.familytreedna.com/learn : accessed 31 May 2014).
- See ISOGG Wiki (http://www.isogg.org/wiki), “Mitochondrial DNA tests,” rev. 30 Mar 2014. ↩
- Virginia Department of Health, Certificate of Death, state file no. 95-011808, Opal Robertson Cottrell (1995); Division of Vital Records, Richmond. ↩
- Wikipedia (http://www.wikipedia.com), “Haplogroup H (mtDNA),” rev. 14 May 2014. ↩
- ISOGG Wiki (http://www.isogg.org/wiki), “Heteroplasmy,” rev. 31 Mar 2014. ↩
- There are actually two reference sequences now in use, the older Cambridge Reference Sequence, also known as the CRS or rCRS, and the newer Reconstructed Sapiens Reference Sequence, RSRS. See generally Roberta Estes, “The CRS and the RSRS,” DNAeXplained, posted 15 July 2012(http://dna-explained.com : accessed 31 May 2014). ↩
- ISOGG Wiki (http://www.isogg.org/wiki), “Genetic distance,” rev. 15 May 2014. ↩



Wow…”nobody ever said this would be easy” is right!
My understanding is that mtDNA is the most consistently stable DNA data of all, at least for genetic genealogy.
Sometimes it seems the more I know the less I realize I know…good timing for a Family History and DNA conference!
Just the place for us all to have a chance to learn some, Mary!
Hi Judy,
To confuse the matter even more, FTDNA uses genetic distance in their yDNA and mtDNA to say how many markers are different between 2 people. 23andMe uses the term to describe the number of cMs two people share on their atDNA.
How does a heteroplasmy occur ?
Nobody appears to be entirely sure. As scientists noted in an article published online: “very little is known about the origin, genetics, and phenotypic effects of heteroplasmic mtDNA mutations.” See Douglas C. Wallace and Dimitra Chalkia, “Mitochondrial DNA Genetics and the Heteroplasmy Conundrum in Evolution and Disease,” Cold Spring Harbor Perspectives in Biology (2013).
Judy, Thank you for this article. I have two brothers who have the same issue – genetic distance of 2 with mtDNA but everything lining up as full brothers with atDNA testing. Someone sent me a link to this article from a face book group to help with explaining how full brothers would not have identical mitochondrial DNA. Unfortunately they are each other’s only matches. Never mind. This was a great article that has helped me understand better.